By Dr Fiona McMillan
A team of researchers at the University of Queensland Diamantina Institute has developed a new method that is set to significantly improve the way we study the human gut microbiome and, by extension, advance our understanding of how the genetics of these microbes affect human health.
Recent years have seen a remarkable increase in evidence pointing toward the importance of the gut microbiome in human health. For example, there is evidence that gut microbes have a profound effect on the development of the human immune system and influence our susceptibility to immune-related diseases. Indeed, it is becoming increasingly clear that your health is influenced by a complex interplay between your genetics, your environment, your lifestyle and your gut microbes. This complexity further increases when you take into consideration the sheer number of microbes in the gut: more than 10 billion a teaspoon. Moreover, not only do the types of gut microbes vary from person to person, but they can also vary significantly within the same person depending on which part of the gut they reside. For example, the microbial communities that reside in your small intestine are very different from those found in the colon.
So how do we find out precisely which microbes drive or mitigate which diseases? It’s an important question, because it could lead to the discovery of new treatments and preventatives for a wide range of debilitating conditions. One of the most powerful approaches is to take samples containing gut microbes so that they can be grown, or ‘cultured’, in the laboratory. Once cultured, the bacteria can be studied to determine how they interact with human cells. In addition, analysis of the bacterial genomes can help us understand which microbial genes affect the development of certain diseases.
This approach sounds straightforward but, in fact, there are major technical challenges preventing both the culturing and genetic analysis of many gut microbes. Consequently, the vast majority of gut microbes remain uncultured and we know very little about most of the human gut microbiome.
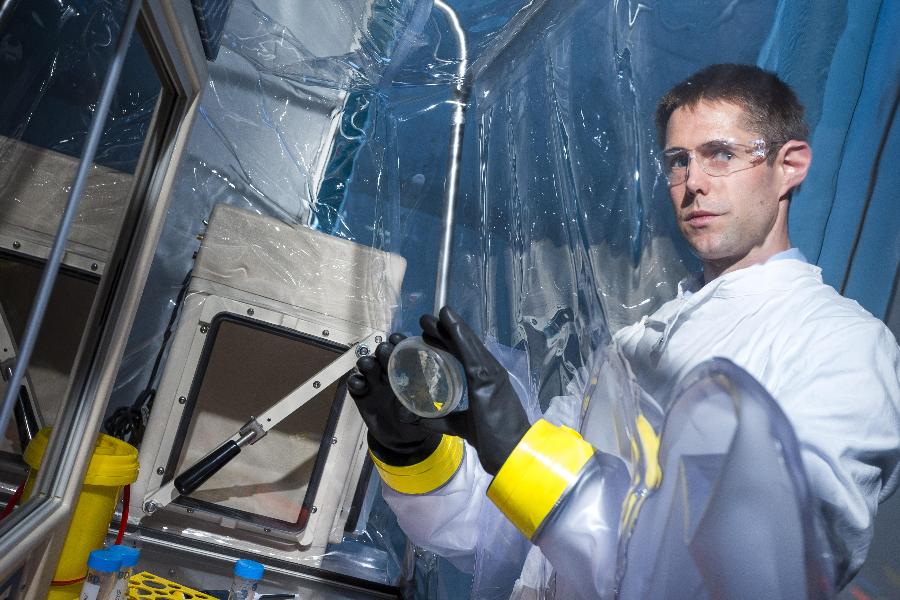
Dr Páraic Ó Cuív is a research fellow at the University of Queensland Diamantina Institute. He explains that the vast majority of gut microbes have evolved a finely tuned preference for growth conditions inside the gut, and replicating these conditions in a laboratory setting can be quite difficult. Furthermore, some gut microbes behave like fast growing “weeds” that outgrow the more interesting microbes in the culture, making them hard to find.
“And for those that are cultured,” he adds, “we lack the techniques to fully characterise them.”
Traditionally, one of the best ways to discover what a particular gene does is to inactivate it and subsequent changes in cell behaviour provide clues as to what a particular gene’s function(s) may be. This approach is called a ‘knockout’. If done systematically, knockouts can build an important picture of a microbe’s functional potential. In fact, this is the approach that was used to successfully study the genome of the bacterium Escherichia coli and why we now know so much about it.
“But the vast majority of gut bacteria are only distantly related to E. coli,” says Ó Cuív. “Many of the methods used to characterise E. coli won’t work for these bacteria.”
This is because, with some notable exceptions, there are few cultured gut bacteria known to be amenable to genetic manipulation. Furthermore, many gut bacteria don’t accept foreign DNA as easily as E. coli using standard laboratory techniques and that makes genetic characterisation incredibly challenging.
Yet, Ó Cuív and his colleague Professor Mark Morrison knew that gut bacteria were capable of sharing DNA with one another, which meant there had to be a way in. It wasn’t a door they were looking for, it was a bridge.
“It’s called microbial conjugation,” says Ó Cuív. “It’s a natural mechanism by which bacteria build a bridge between each other so that they can transfer DNA.”
Interestingly, it turns out that when it comes to microbial conjugation, bacteria can be quite promiscuous and specific E. coli strains can build bridges and move DNA into a large range of other bacteria. Moreover, they can also do the same with archaea, fungi and even animal cells.
In order to determine the extent of genetically modifiable bacteria in the gut, Ó Cuív and his colleagues wanted to see if they could get foreign DNA into gut bacteria by using E. coli as a stepping stone. In other words, foreign DNA is first transferred into E.coli which then forms bridges with the gut bacteria and passes it along. In order to have a clear indication of whether or not the approach worked, the foreign DNA they used included a gene that confers resistance to a particular antibiotic. This meant that any gut bacterium that successfully acquired the foreign DNA would be able to grow in the presence of that antibiotic. It worked even better than they expected.
“It’s incredibly efficient and a very rapid process,” says Ó Cuív. “We have shown that these bacteria are capable of receiving foreign DNA from E. coli, suggesting that they are amenable to genetic characterisation.”
The method, which was recently published in the journal Scientific Reports, also improves the ability to culture bacteria of interest by enabling the researchers to restrict the growth of unwanted, nutrient-greedy bacteria.
The UQDI research group has now established the world’s largest culture collection of gut bacteria that are amenable to genetic study, explains Ó Cuív.
“The next goal is to construct specific gene knockouts to better understand the bacteria,” he adds. “We are now beginning to screen them for specific functional activities so that we can prioritise them for further study.”
For example, this process could help identify the genes responsible for the anti-inflammatory effects associated with certain gut bacteria. This, in turn, could facilitate the development of new anti-inflammatory drugs.
The approach is also allowing the research team to study where different bacteria grow best and how they behave in different regions of the GI tract. This is important because gut bacteria can have vastly different effects on their hosts depending on which part of the gut they inhabit. For instance, the chemical signals they send out can change, affecting which human genes are turned on and off in response.
“If the bacterium goes to the wrong place it may affect your health and well-being,” says Ó Cuív.
Take the case of the gut bacterium Clostridium ramosum, which is associated with increased obesity in humans. Recent research shows that C. ramosum causes mice on a high fat diet to be much more susceptible to obesity. Interestingly, this bacterium triggers very different effects on the host depending on where it is located in the small intestine.
The UQDI research team have now used their new method to genetically tag gut bacteria including C. ramosum, making it easier to monitor their location and behaviour. In much the same way that ecologists tag and track animals in the wild, this allows the researchers to learn more about which gut environments a particular microbe prefers, which environments cause stress, and what might cause it to move from one place to another.
Ó Cuív points out that their approach is readily adaptable and could be used for the study of microbes elsewhere in the body, and anywhere else microbial diversity is of interest, such as in agricultural or ecological research. Consequently, the UQDI team is very excited about the impact this approach will have not only on the study of the human gut microbiome, but also on our broader understanding of the microbial world around us.
Media: Kate Templeman, k.templeman@uq.edu.au +61 7 3443 7027, +61 (0)409 916 801.